Tandem mass spectrometry: Difference between revisions
CSV import |
CSV import |
||
| Line 22: | Line 22: | ||
{{stub}} | {{stub}} | ||
<gallery> | |||
File:Q-TOF.jpg|Q-TOF Mass Spectrometer | |||
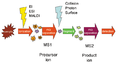
File:MS_MS.png|MS/MS Process | |||
File:Triple_quadripole.png|Triple Quadrupole Mass Spectrometer | |||
File:LTQ_Trap_And_Dynodes_2.jpg|LTQ Trap and Dynodes | |||
File:Isobaric_labeling.png|Isobaric Labeling in Mass Spectrometry | |||
File:PeptideMSMS.jpg|Peptide Fragmentation in MS/MS | |||
</gallery> | |||
Latest revision as of 04:36, 18 February 2025
Tandem mass spectrometry (also known as MS/MS or MS2) is a method used in analytical chemistry to identify the quantity and type of individual chemical compounds in a sample. It involves multiple steps of mass spectrometry and some form of fragmentation.
Overview[edit]
In a tandem mass spectrometry process, ions are formed in the ion source and are then subjected to a first stage of mass spectrometry to separate them by their mass-to-charge ratio. The selected ions are then fragmented and the fragments are subjected to a second stage of mass spectrometry to further separate them. The resulting spectrum is then used to identify the original molecule.
Applications[edit]
Tandem mass spectrometry is used in many different fields, including biochemistry, pharmacology, and environmental science. It is particularly useful in proteomics, where it is used to identify proteins and to determine their sequences. It is also used in metabolomics to identify small molecules.
See also[edit]
References[edit]
<references />